|
Ggplot(rawData, aes(koerpergewicht, groesse, color = factor(cluster$x))) + Labels = c("1 schwer", "2 leicht","3 Zwischengruppe"), Geom_point() + labs(title = paste(nlevels(factor(colors))))+geom_point(size=8)+geom_text(aes(label=position),vjust=-1.5)+scale_color_manual(name = "Gruppe", Ggplot(rawData, aes(koerpergewicht, groesse, color = factor(data$gruppe))) + PlotOutput("plot2", hover = "plot_hover"), I could imagine you dont want to specify the code for plot/table twice in shiny + rmarkdown, but i guess you would have to choose between both options and the first one is probably cleaner. Ui<-fluidPage(plotOutput("plot1", hover = "plot_hover"), I have already tried: options(width = 800)ĭata<-read.csv2('data.csv',header=T,sep=",") I want to get rid of the scrollbars and show the 2 figures with full width and height without scrollbars. It works fine but the output is in a really small area with scrollbars. An extra addition could be that first there are radiobuttons on top to chose which colum you would like to filter on (eg $test), under it a drop down menu to chose the value of the filter and under that the histogram.I want to generate a R markdown page with an interactive shiny app. Since it is not really possible to run code chunks (with dynamic renderplots) separately, it takes a lot of time to debug the script. It is still a draft script and there is room for (esthetical) improvements. This article will show you how to write an R Markdown report. You write the report in markdown, and then launch it as an app with the click of a button. An interactive document is an R Markdown file that contains Shiny widgets and outputs. Names(df_aggr_Result) <- "amount of samples"īarplot(df_aggr_Result, probability = TRUE, JInteractive documents are a new way to build Shiny apps. #Calculate the amount of Result-samples each hour:ĭf2 <- as.ame(hour(hms(df1_filtered$ResultTime)))ĭf_aggr_Result <- aggregate(df2, by=list(df2$`hour(hms(df1$ResultTime))`), FUN = length) SelectInput("Validation source", label="Who/What performed the validation:",Ĭhoices = df1$ValidationUser, selected = "~SYSValDaemon~")ĭf1_filtered <- df1 I tried already some codes, but the aggregate seems the give an error, since It cannot find any rows? library(lubridate) 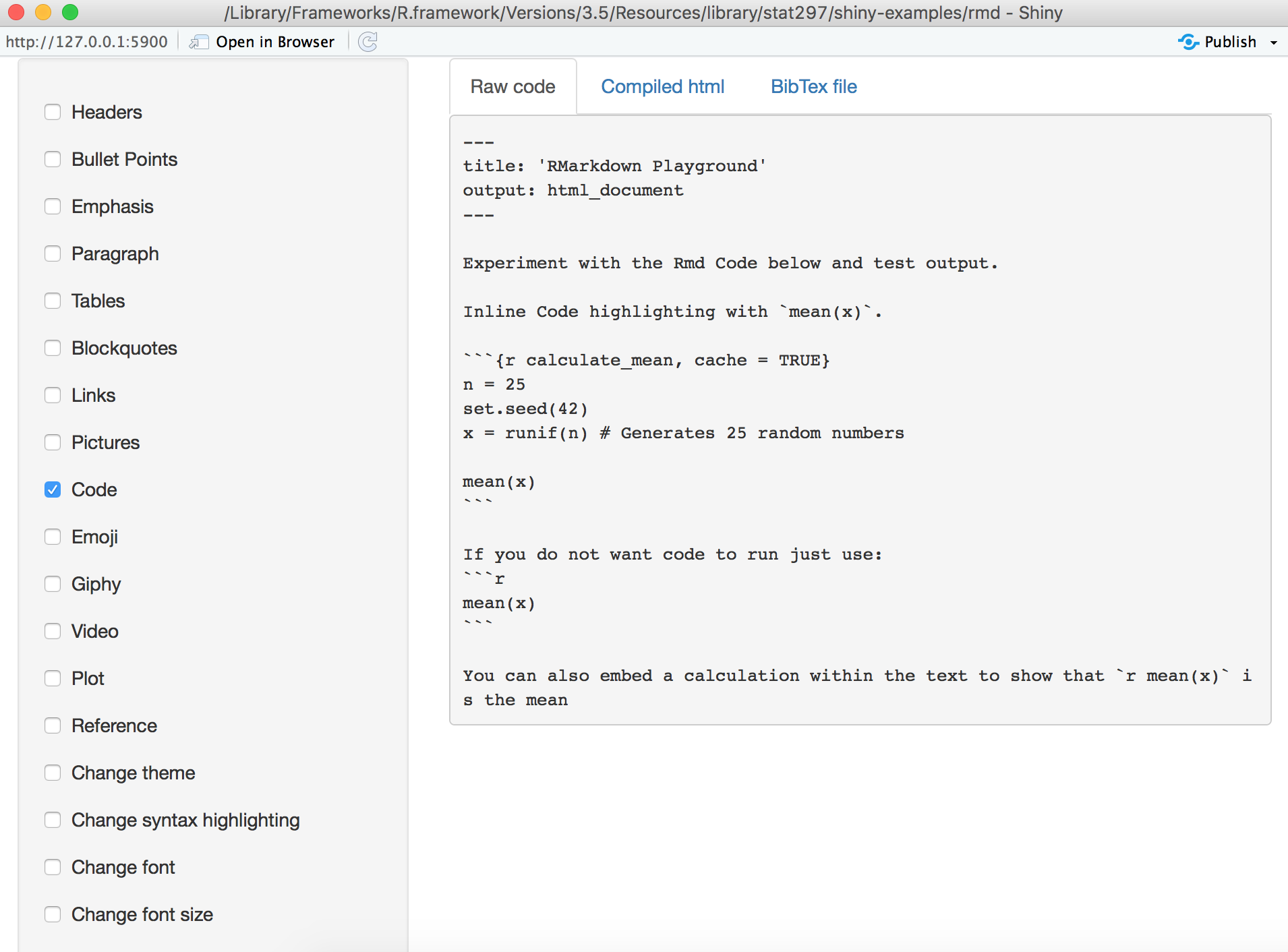
Labs(title="amount of results over the day",x="hours", y = "amount of samples")+īut now before making the histogram, I should create a filter on the data, based on the selectInput variable. Mapping = aes(x = df_aggr_Result$hour, y = df_aggr_Result$`amount of samples`)) + Theme(plot.title = element_text(hjust = 0.5)) + 
Labs(title="amount of first-scan times over the day",x="hours", y = "amount of samples") + Geom_bar(stat="identity", boundary = 0, color="blue", fill="lightblue") + 
what I would like to have is that I call a bask script and the dynamic report is created and opened in a browser. Now I would like to use an interactive report (i.e. This uses rmarkdown::render() and works nicely. Mapping = aes(x = df_aggr_First_scan$hour, y = df_aggr_First_scan$`amount of samples`)) + This (static) report is, via a bash script which calls R, created as a pdf or an html. I used following code to make a barplot of the amount of samples each hour of all the samples in r markdown: Another example is show the histogram, representing the amount of sample each hour of $test IGF. 
Each example below contains a link to the source code within the dashboard. Dashboards Combine R Markdown with the flexdashboard package to quickly assemble R components into administrative dashboards. eg: show the histogram, representing the amount of sample each hour for $Source(location) A. Shiny Shiny components and htmlwidgets will work in any HTML based output, such as a file, slide show or dashboard. Now the data is kind of prepared I would like to make a histogram/barplot to give the amount of samples over time (1 hour interval) and to use a dropdown menu to select an extra filter. I hope this give the same result on the Shiny app. #The difference between the ResultTime and the FirstScanTime is the turn around time:ĭf1$TS_start <- paste(df1$FirstScanDate, df1$FirstScanTime)ĭf1$TS_end <- paste(df1$ResultDate, df1$ResultTime)ĭf1$TAT <- difftime(df1$TS_end,df1$TS_start,units = "mins") #Create a subset of the data to remove/exclude the unnecessary columns: Server_data <- lim(file="PostCheckAnalysisRoche.txt", header = TRUE, na.strings=c(""," ","NA")) Setwd("G:/My Drive/Traineeship Advanced Bachelor of Bioinformatics 2022/Internship 2022-2023/internship documents/") I use the code: #First the exported file from the Infinity server needs to be in the correct folder and read in:
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
